-
17
11 Month

The 2024 Nobel Prize in Physiology or Medicine: microRNA Shines Brilliantly!
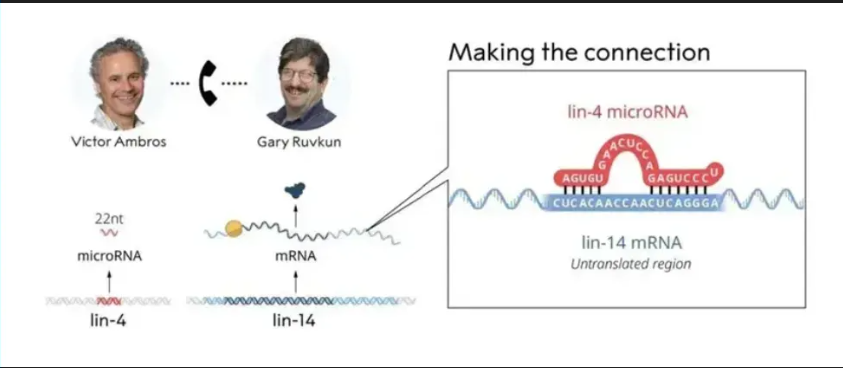
On October 7th, 2024, the Karolinska Institute in Sweden announced that the 2024 Nobel Prize in Physiology or Medicine was awarded to two American scientists, Victor Ambros and Gary Ruvkun, in recognition of their discovery of microRNA (microribonucleic acid) and its role in post-transcriptional gene regulation.

As early as 1989, scientist Victor Ambros discovered a gene named lin-4 in the nematode (C. elegans), which could suppress the expression of another gene, lin-14. This discovery hinted that lin-4 might express a regulatory protein, but subsequent research revealed that the product of lin-4 was actually an RNA molecule with a length of only 22 nucleotides. In 1993, Victor's students Rosalind Lee and Phonda Feinbaum successfully cloned the lin-4 gene and confirmed that its product was a non-coding RNA, namely microRNA.
-
27
09 Month

Application Guide for Neohalo Bioscience Trial Kits
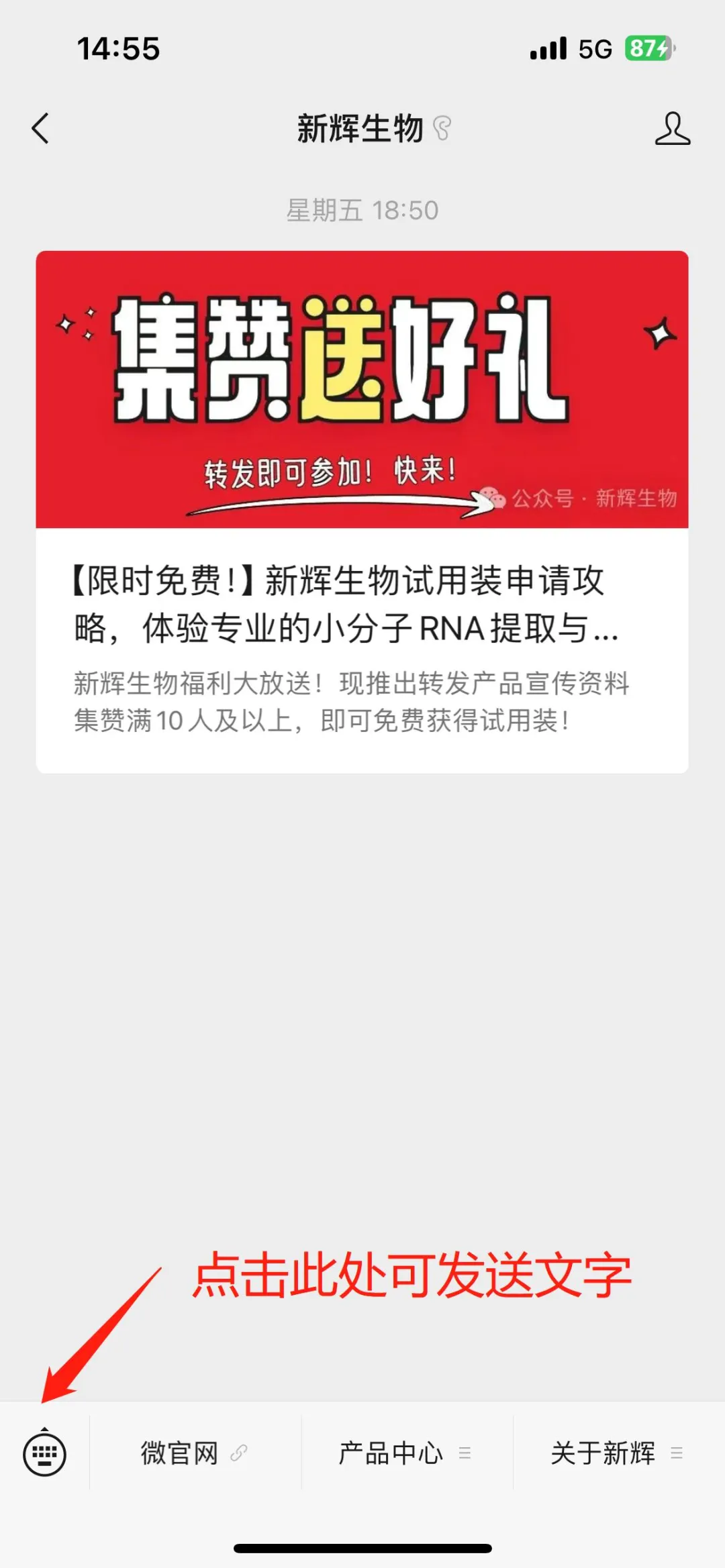
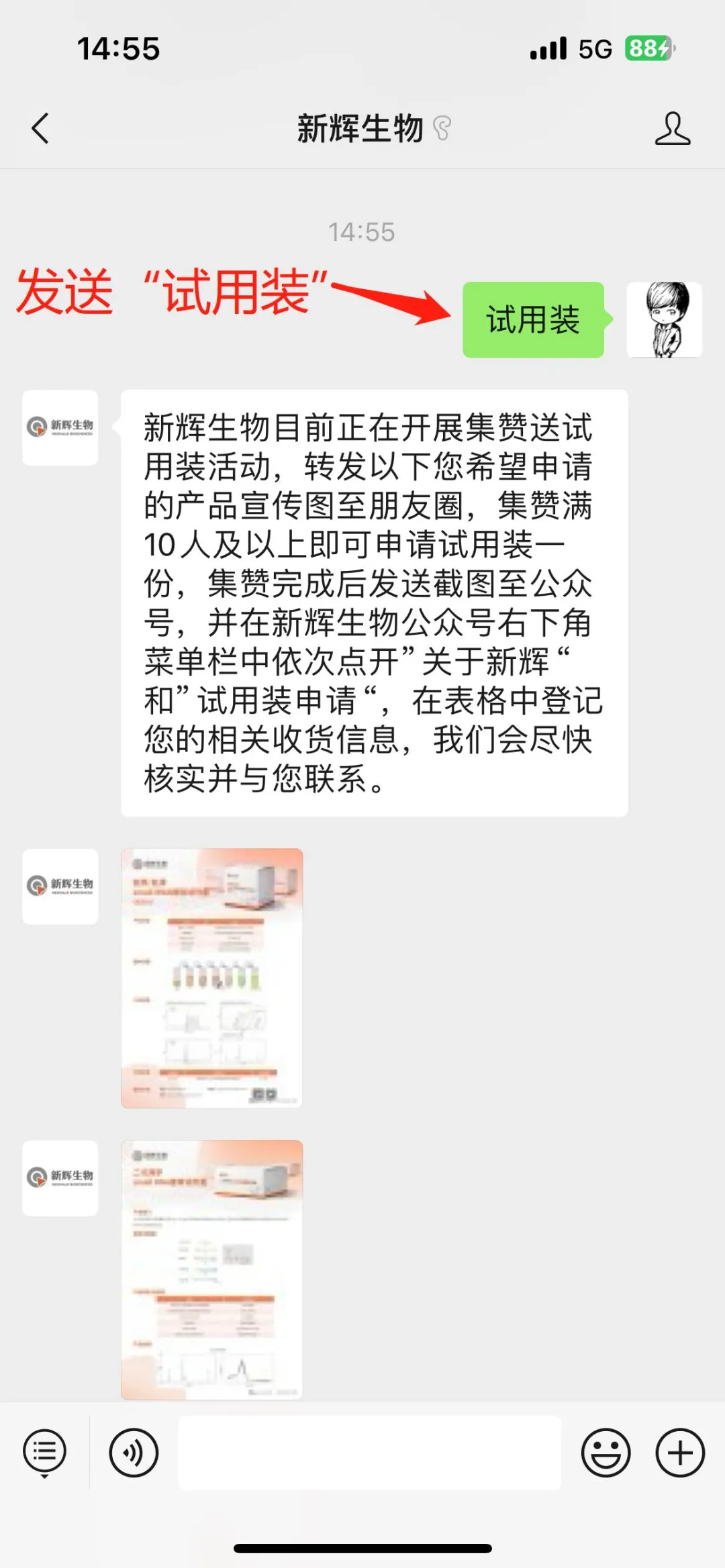
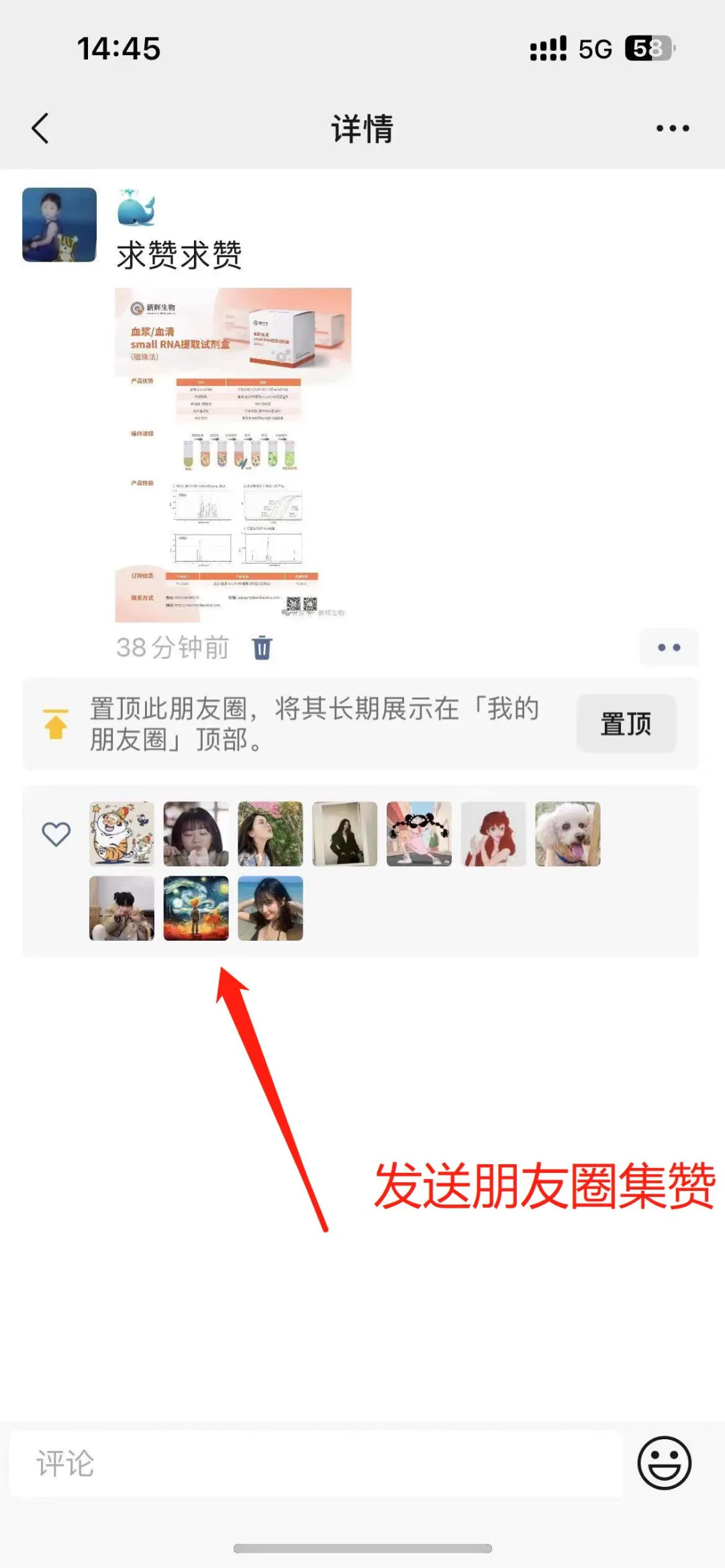
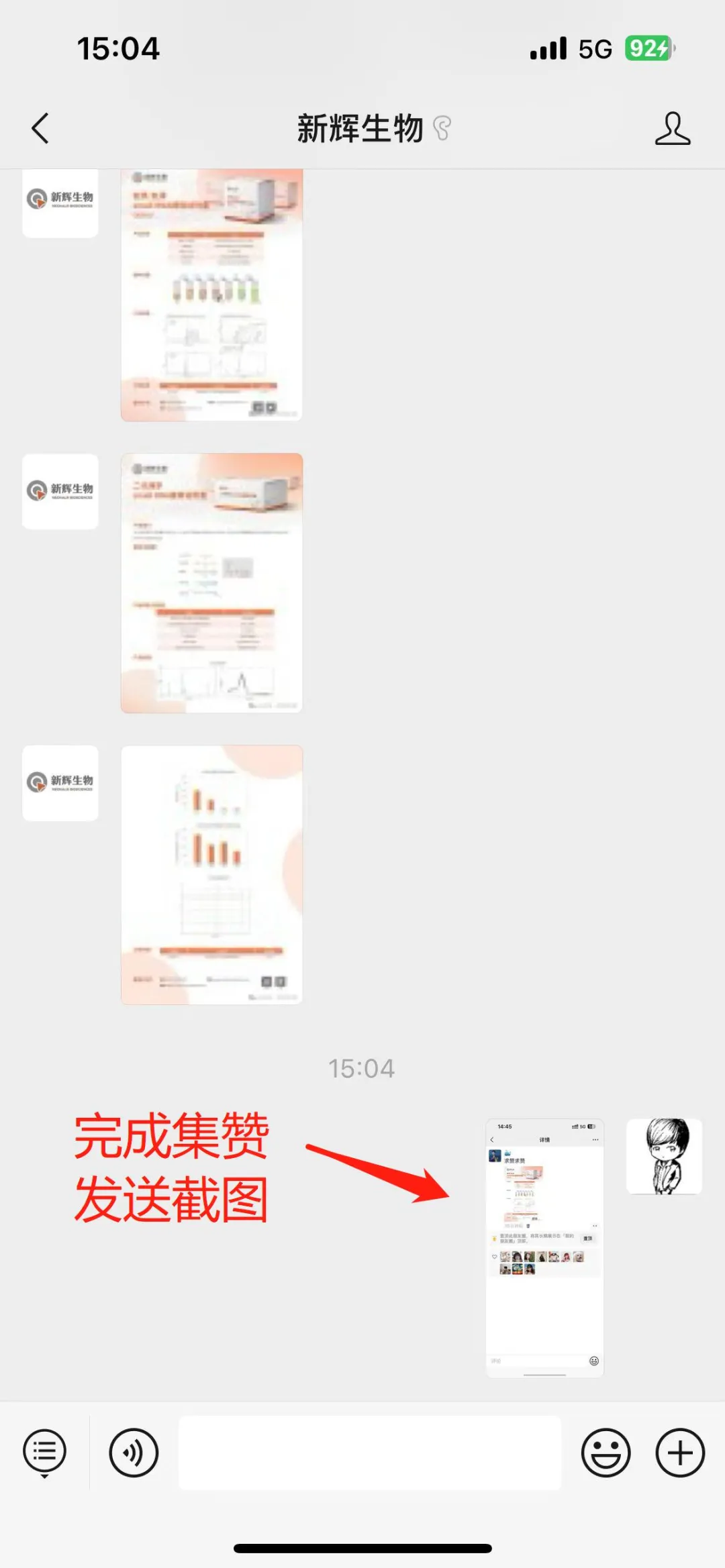
1.Obtain Forwarding Materials
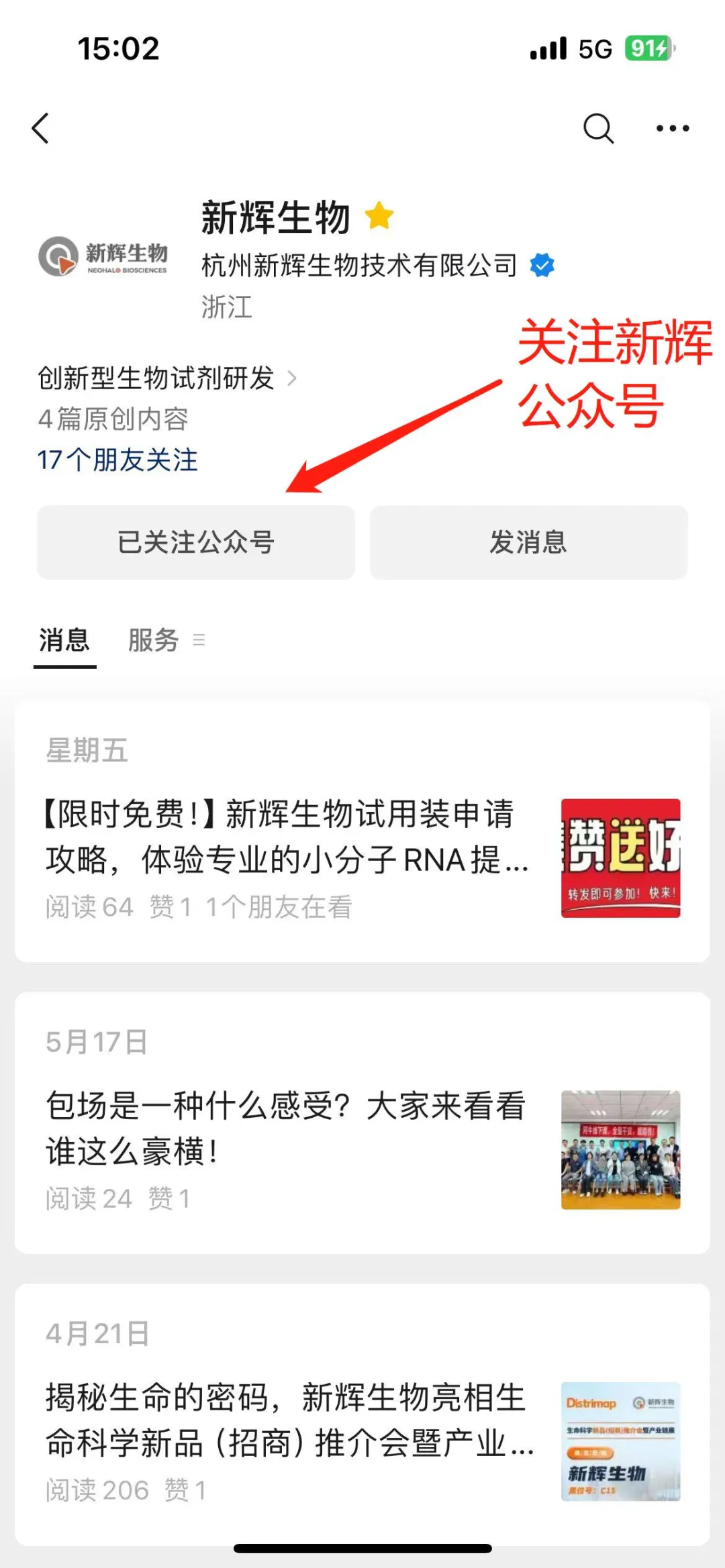
1.Firstly, new friends need to follow our official WeChat account first. Directly search for "XinHui Biology" in WeChat and click to follow.
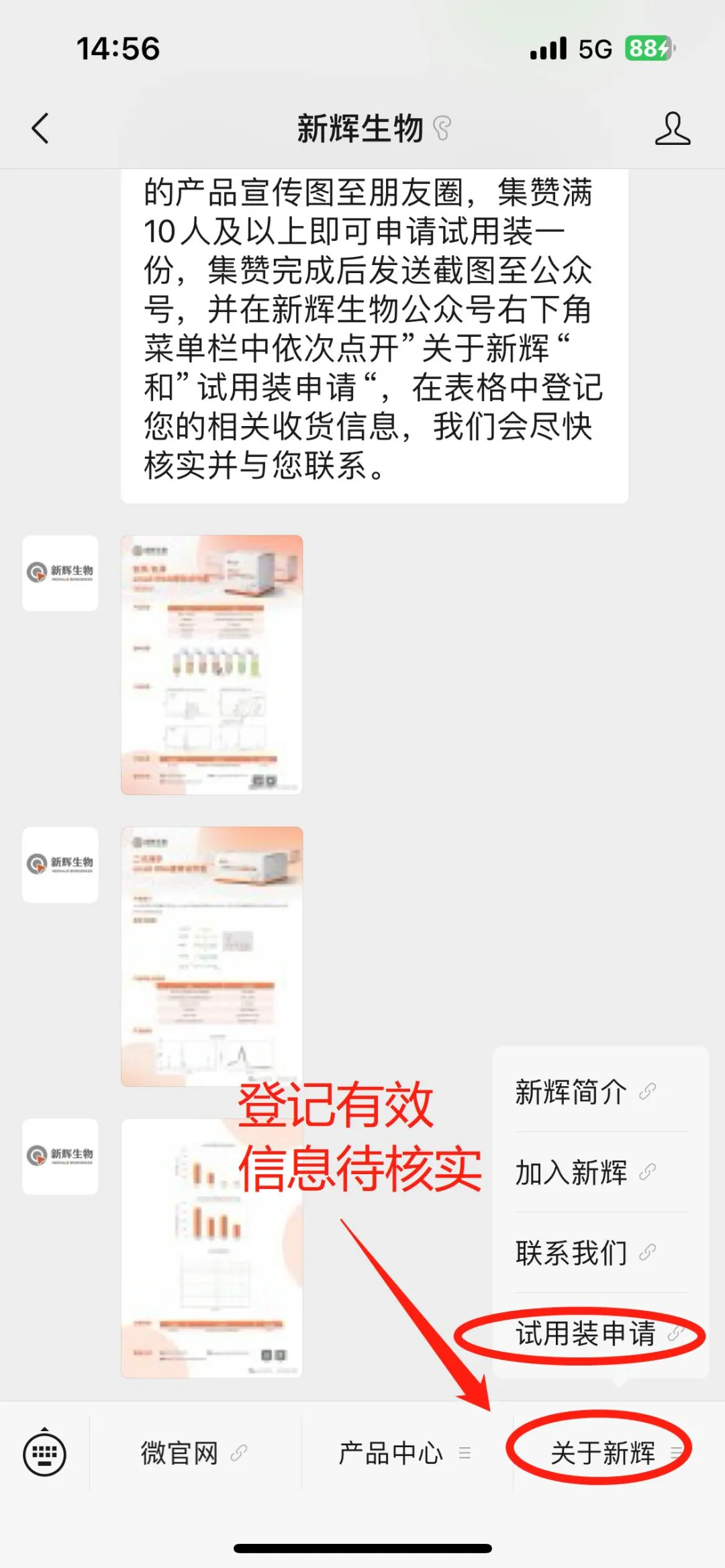
3. Self-registration on the Official WeChat Account 1. After sending the screenshot of completing the like collection, click on "About Neohalo Bioscience" and then "Application for Trial Kits" in the sub-menu bar in turn at the bottom right corner of the Neohalo Bioscience official WeChat account. Fill in the valid information truthfully so that we can review it later and get in touch with you.
4. Activity Summary Dear friends, as long as you follow the steps in the guide one by one, I believe you will be able to use the desired trial kits soon! I'd like to tell you secretly that if your research direction matches our products very well, we sincerely invite you to get in touch with us by phone to further discuss cooperation matters. We will give priority to your application for the kits.
Activity Rules: 1. Application Frequency: Each customer has only one free application opportunity, and only one type of trial kit product can be obtained each time. If you exceed the limit, you need to purchase it by yourself.
2. To avoid reagent waste, we will review your application materials one by one. To avoid delaying your time, please fill in the registration materials truthfully so that we can get in touch with you later.
3. The available stock of trial kits is limited. First come, first served.